The main goal of the tool is to use these features in toxicity modelling. Recently a set of generic chemical functionalities called ToxPrint chemotypes that describe molecule substructure and reaction features and atom and bond properties was defined within the ToxPrint program. Another widely used set of substructures used to identify potentially reactive or promiscuous molecule has been defined by Eli Lilly scientists based on their experience with internal screening campaigns.
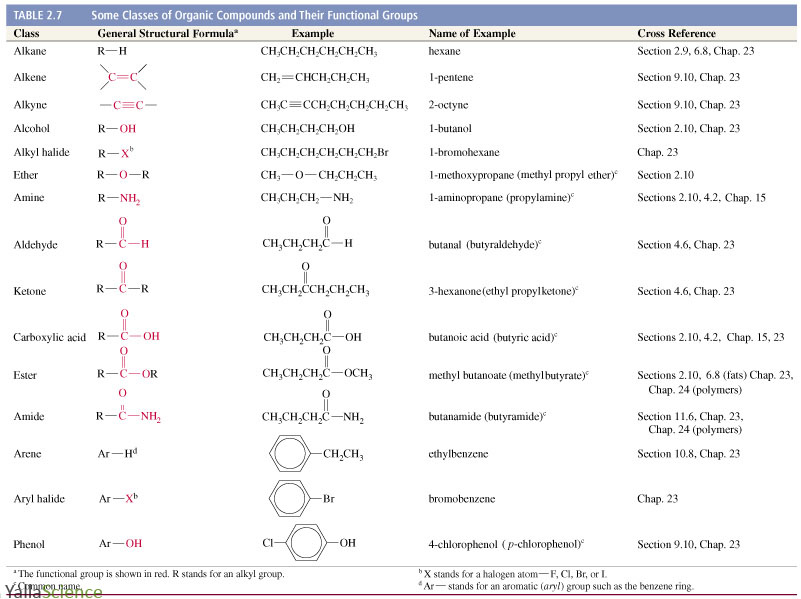
The patterns include PAINS filters that identify frequent hitters interfering with biochemical screens as well as some other substructures. The well-known ZINC database and related web-based software suite stores about 500, so called, chemical patterns, that speed-up substructure searches and allow estimation of molecule reactivity. These substructure features are used to develop QSAR models for prediction of toxicity and various molecular physicochemical properties. 5-membered aromatic ring with 1 heteroatom). This list includes also numerous heterocyclic rings and general structural patterns (i.e. Recently an extended version of the program containing 583 manually curated functionalities encoded as SMARTS was published. One of the first software tools to identify FGs was the checkmol program written by Haider that was able to identify about 200 FGs. The fragments are generally strongly overlapping and are generated for all parts of a molecule without considering their potential chemical role. Although such fragment descriptors are very useful, they do not provide description of functional groups. Examples of such descriptors are linear or atom centered fragments, topological torsions, pharmacophoric triplets and many others. In this approach the substructure descriptors are generated by extracting groups of atoms from a molecule using a predefined algorithm. Various substructure features are often used in cheminformatics in connection with machine learning to develop models to predict biological activity or properties of molecules. describing a rule-based definition of chemical classes to classify compounds into classes or the ClassyFire software developed in the Wishart’s group allowing chemists to perform large-scale automated chemical classification based on a structure-based chemical taxonomy consisting of over 4800 categories. An example of this type of publications is work by Bobach et al. The majority of theoretical studies are utilizing FGs as a basis of chemical ontologies, where FGs are “keys” that are used to hierarchically classify molecules into categories. There is, however, surprisingly little attention paid to the study of functional groups from the cheminformatics point of view. A well known example is the classical book series “Chemistry of functional groups” describing various classes of organic molecules consisting of over 100 volumes. Numerous scientific papers and books focus on properties and reactivity of various FGs. The study of common FGs forms substantial part of basic organic chemistry curriculum. The concept of functional groups (FGs)-sets of connected atoms that determine properties and reactivity of parent molecule, forms a cornerstone of organic chemistry, medicinal chemistry, toxicity assessment, spectroscopy and, last but not least, also chemical nomenclature. The new method allows the analysis of functional groups in large chemical databases in a way that was not possible using previous approaches. The algorithm is relatively simple and full details with examples are provided, therefore implementation in any cheminformatics toolkit should be relatively easy. ConclusionsĪ new algorithm to identify all functional groups in organic molecules is presented. The procedure is illustrated by extracting functional groups from the bioactive portion of the ChEMBL database, resulting in identification of 3080 unique functional groups. ResultsĪn algorithm to identify functional groups in a molecule based on iterative marching through its atoms is described. The algorithm presented in this article is an attempt to solve this scientific challenge. We are not aware of any program that can identify all functional groups in a molecule automatically.
All current software systems to identify functional groups are based on a predefined list of substructures. The concept of functional groups forms a basis of organic chemistry, medicinal chemistry, toxicity assessment, spectroscopy and also chemical nomenclature.
